Mars Documentation¶
Mars is a tensor-based unified framework for large-scale data computation which scales numpy, pandas, scikit-learn and many other libraries.
Architecture Overview¶
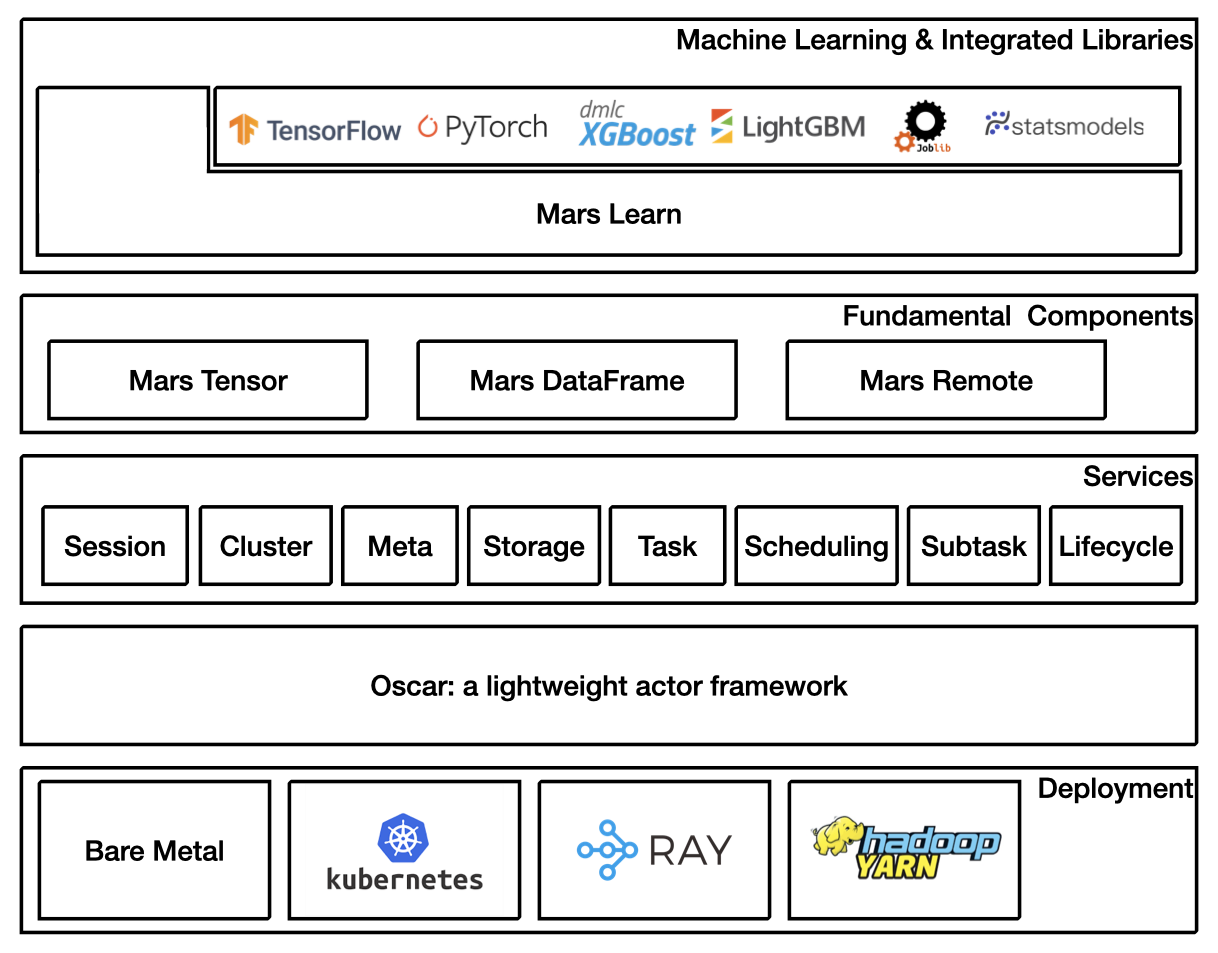
Getting Started¶
Starting a new runtime locally via:
>>> import mars
>>> mars.new_session()
Or connecting to a Mars cluster which is already initialized.
>>> import mars
>>> mars.new_session('http://<web_ip>:<ui_port>')
Mars tensor¶
Mars tensor provides a familiar interface like Numpy.
Numpy |
Mars tensor |
import numpy as np
N = 200_000_000
a = np.random.uniform(-1, 1, size=(N, 2))
print((np.linalg.norm(a, axis=1) < 1)
.sum() * 4 / N)
|
import mars.tensor as mt
N = 200_000_000
a = mt.random.uniform(-1, 1, size=(N, 2))
print(((mt.linalg.norm(a, axis=1) < 1)
.sum() * 4 / N).execute())
|
3.14174502
CPU times: user 11.6 s, sys: 8.22 s,
total: 19.9 s
Wall time: 22.5 s
|
3.14161908
CPU times: user 966 ms, sys: 544 ms,
total: 1.51 s
Wall time: 3.77 s
|
Mars can leverage multiple cores, even on a laptop, and could be even faster for a distributed setting.
Mars dataframe¶
Mars DataFrame provides a familiar interface like pandas.
Pandas |
Mars DataFrame |
import numpy as np
import pandas as pd
df = pd.DataFrame(
np.random.rand(100000000, 4),
columns=list('abcd'))
print(df.sum())
|
import mars.tensor as mt
import mars.dataframe as md
df = md.DataFrame(
mt.random.rand(100000000, 4),
columns=list('abcd'))
print(df.sum().execute())
|
CPU times: user 10.9 s, sys: 2.69 s,
total: 13.6 s
Wall time: 11 s
|
CPU times: user 1.21 s, sys: 212 ms,
total: 1.42 s
Wall time: 2.75 s
|
Mars learn¶
Mars learn provides a familiar interface like scikit-learn.
Scikit-learn |
Mars learn |
from sklearn.datasets import make_blobs
from sklearn.decomposition import PCA
X, y = make_blobs(
n_samples=100000000, n_features=3,
centers=[[3, 3, 3], [0, 0, 0],
[1, 1, 1], [2, 2, 2]],
cluster_std=[0.2, 0.1, 0.2, 0.2],
random_state=9)
pca = PCA(n_components=3)
pca.fit(X)
print(pca.explained_variance_ratio_)
print(pca.explained_variance_)
|
from mars.learn.datasets import make_blobs
from mars.learn.decomposition import PCA
X, y = make_blobs(
n_samples=100000000, n_features=3,
centers=[[3, 3, 3], [0, 0, 0],
[1, 1, 1], [2, 2, 2]],
cluster_std=[0.2, 0.1, 0.2, 0.2],
random_state=9)
pca = PCA(n_components=3)
pca.fit(X)
print(pca.explained_variance_ratio_)
print(pca.explained_variance_)
|
Mars learn has also integrated many libraries, including tensorflow, xgboost, lightgbm, joblib and statsmodels.
Mars remote¶
Mars remote allows users to execute functions in parallel.
import numpy as np
def calc_chunk(n, i):
rs = np.random.RandomState(i)
a = rs.uniform(-1, 1, size=(n, 2))
d = np.linalg.norm(a, axis=1)
return (d < 1).sum()
def calc_pi(fs, N):
return sum(fs) * 4 / N
N = 200_000_000
n = 10_000_000
fs = [calc_chunk(n, i)
for i in range(N // n)]
pi = calc_pi(fs, N)
print(pi)
|
import numpy as np
import mars.remote as mr
def calc_chunk(n, i):
rs = np.random.RandomState(i)
a = rs.uniform(-1, 1, size=(n, 2))
d = np.linalg.norm(a, axis=1)
return (d < 1).sum()
def calc_pi(fs, N):
return sum(fs) * 4 / N
N = 200_000_000
n = 10_000_000
fs = [mr.spawn(calc_chunk, args=(n, i))
for i in range(N // n)]
pi = mr.spawn(calc_pi, args=(fs, N))
print(pi.execute().fetch())
|
3.1416312
CPU times: user 32.2 s, sys: 4.86 s,
total: 37.1 s
Wall time: 12.4 s
|
3.1416312
CPU times: user 616 ms, sys: 307 ms,
total: 923 ms
Wall time: 3.99 s
|
Easy to scale in and scale out¶
Mars can scale in to a single machine, and scale out to a cluster with hundreds of machines. Both the local and distributed version share the same piece of code, it’s fairly simple to migrate from a single machine to a cluster to process more data or gain a better performance.
Mars can run in a few ways: